Autophagy is a catabolic transport route conserved among all eukaryotes that allow the degradation of large portions of the cytoplasm, protein aggregates, excess or damaged organelles and invading pathogens.

guest appointment section Molecular Cell Biology,
1) The regulation and mechanism of autophagy (in yeast Saccharomyces cerevisiae) 2) Autophagy-pathogen interactions

Postdoctoral fellows:
Franziska Kriegenburg (2014 – 2018)
Jana Sanchez-Wandelmer (2012 – 2016)
Henning Arlt (2014-2016)
Muriel Mari (2006 – 2015)
Eduardo Cebollero (2008-2012)
Kristy Horan (2012)
Iryna Monastyrska (2007–2010)
Adabella van der Zand (2009-2010, visiting postdoctoral fellow)
PhD students:
Nienke van Beek (2014-2019)
Yingying Cong (2015–2019)
Qianliang Yuan (2017–2019)
Xingdong Zhou (2016-2017)
Rodrigo Guimaraes (2013-2017)
Xingdong Zhou (2016-2017)
Andri Fraenkl (2013-2016)
Susana Abreu (2012-2016)
Leticia Lemus (2015, visiting PhD student)
Joanna Liiv (2014, visiting PhD student)
Ester Rieter (2008-2012)
Shan Shan Wang (2012, visiting PhD student)
Mustafa Ulasli (2007-2011)
Aniek van der Vaart (2006-2010)
Nian Liu (2008, visiting PhD student)
Daria Romanyuk (2008, visiting PhD student)
Technicians:
Despina Xanthakis (2012–2015)
Janice Griffith (2005–2012)
Master students:
Fleur Broek (2016)
Wouter Huiting (2015)
Alex Jones (2014)
Philip Vkovski (2013)
Mareike Nolte (2012)
Stephanie Keppes (2010–2011)
Ana Maria Guzman-Prieto (2010)
Dorothee van Breevoort (2008-2009)
Fabian Finke (2007-2008)
Ester Rieter (2007-2008)
Jiang Jieqing (2007-2008)
Lakshmi Krishnappa (2007-2008)
Technician trainees:
Rianne Grond (2016)
Kerst-Jan Hijlkema (2015-2016)
Elena Iskandarani (2012)
Remko Goossens (2010)
Marinke van Oorschot (2009-2010)
Nina Bakker (2009-2010)
Wresti Listu Anggayasti (2009)
Tineke Hoefnagel (2008)
Tessa Hoogenhuijzen (2007)
Jan Hazeleger (2006)
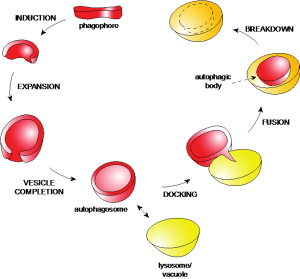
Autophagy is a catabolic process highly conserved among eukaryotes, which is involved in the degradation of long-lived proteins, aberrant complexes and aggregates, dysfunctional and superfluous organelles, and intracellular pathogens. Structures targeted to destruction are sequestered by double-membrane vesicles called autophagosomes and delivered to lysosomes for turnover. The resulting metabolites are reused by the cell as either an energy source or building blocks for the synthesis of new macromolecules.
Autophagy is essential to maintain cellular and organismal homeostasis because involved in the adaptation to stresses, quality control, metabolism regulation, cell development and differentiation, stemness maintenance, type II program cell death, tumor suppression and immunity. As a result, the impairment or defect in autophagy leads to severe pathologies such as neurodegeneration, myopathies, chronic inflammations, and some malignancies. Crucially, it has also been shown that autophagy is an effective therapy to prevent or cure diseases, including specific types of tumors, muscular dystrophies, and neurodegenerative disorders. The 2016 Nobel Prize in Medicine or Physiology to Prof. Ohsumi, a leading scientist in the field, underlines the recognized importance of autophagy in medical and life sciences.
The elucidation of the mechanisms involved in activation and regulation of autophagy, but also the identification of the proteins required for recognition and elimination of the various autophagic cargoes, is therefore of primary relevance. Understanding the exact role of autophagy in the various physiological situations and pathological conditions is also crucial. All this knowledge combined is essential to be able to modulate autophagy to the benefit of human health, but also for biotechnological and agricultural applications.
The laboratory has two major research lines. In the first, we aim in unveiling the regulation and molecular mechanism of autophagy, and for these studies, we use yeast Saccharomyces cerevisiae as a model system. As the long-term objective is to understand the exact contribution of autophagy in specific physiological and pathological contexts, the goal of the second research line has been to understand the interaction between autophagy and pathogens, in particular viruses. In doing this, our interests have also moved to the characterization of unconventional, not autophagy-linked, function of the autophagy proteins.
Dissertations supervised by Fulvio M. Reggiori ((co-) promotor or assessor):
2019
2018
Yuan, W. (2016). Origin and growth of peroxisomes in yeast: The molecular mechanism of peroxisome formation in yeast. [Groningen]: University of Groningen.
2012
Rieter, E. (2012). The molecular organization of the phagophore assembly site [Utrecht]: University of Utrecht
2011
Ulasli, M. (2011). Coronavirus replication in host cells [Utrecht]: University of Utrecht
2010
Van der Vaart, A. (2010). Membrane dynamics during autophagy in yeast Saccharomyces cerevisiae [Utrecht]: University of Utrecht
Update your browser to view this website correctly. Update my browser now